Introduction of Dengue Serotyping Real-Time PCR Test
Table of Contents
Dengue is a mosquito (Aedes aegypti)-borne viral disease. It is a major health threat in the tropical and sub-tropical territories of the world, particularly in urban and semi-urban areas. Approximately 50-100 million individuals have been infected with dengue annually (WHO data) and more than 100 countries are at risk of dengue transmission. The following activities are responsible for contributing to fertile breeding areas for the mosquito (Dengue vector)-fast unplanned urbanization and migration of population from rural to urban sectors, climatic changes, no vector control, and poor sanitation services.
There are four serotypes of the Dengue virus. They are Dengue serotype-1 (DENV 1), Dengue serotype-2 (DENV 2), Dengue serotype-3 (DENV 3), and Dengue serotype-4 (DENV 4). DENV-2 and DENV-4 are comparatively highly severe than DENV-1 and DENV-3. Detection of these serotypes is based on the real-time amplification and detection of the dengue virus in a one-step format by Real-Time PCR Test.
Principle of Dengue Serotyping Real-Time PCR Test
The first remarkable increase in the amount of PCR product correlates to the beginning amount of the target template. The higher the starting copy number of the RNA target, the short a significant increase in fluorescence is seen. A significant increase in fluorescence above the baseline value measured during the 3-15 cycles indicates the detection of accumulated PCR product. Usually, the protocol followed is depicted. Real-time PCR is a variation of the standard PCR technique used to quantify messenger RNA (mRNA) in a specimen.
Using sequence-specific primers, the relative number of copies of a particular RNA sequence can be determined. The quantification appears by measuring the amount of amplified product at each stage during the PCR cycle. RNA from genes with higher copy numbers will appear after fewer melting, annealing, and extension PCR cycles. Quantification of amplified product is obtained using fluorescent probes and specialized machines that measure fluorescence while performing temperature changes needed for the PCR cycles.
Test Requirements
For Template (Extracted RNA): For this purpose following are mandatory-
- Nucleic acid (RNA) Extraction kit ( automation e.g. BIOER)
- Automation
- Biosafety cabinet
- Micropipette
- Waste bin
- Centrifuge
- Test specimen (blood)
- Automation cartridge
- Refrigerator
- Ethanol
- Hypochlorite
- Tissue paper
- Other than automation needs-
- Eppendorf tube
- Collection tube
- Manual RNA extraction kit contains-
- Spin column
- Lysis buffer
- Enzyme
- Wash buffer
- Elution buffer
- The Dengue Serotyping PCR test kit contains-
- Dengue Serotyping PCR reaction fluid
- RT-PCR enzyme
- Dengue Serotyping positive quality control
- Negative quality control
- Thermocycler (PCR machine )
- Micropipettes with filter tips
- PCR cabinet
- PCR tubes with cap
- PCR tube holder
Test Procedure of Dengue Serotyping Real-Time PCR Test
- Extract RNA (template) either automation or manually (follow kit manual).
- Prepare master mix (according to the direction of kit manufacturers).
- Add template to a master mix.
- Load PCR tubes in a thermocycler.
- Check the position and program.
- Run the test.
- Wait for completing the run.
- Analyze the graph.
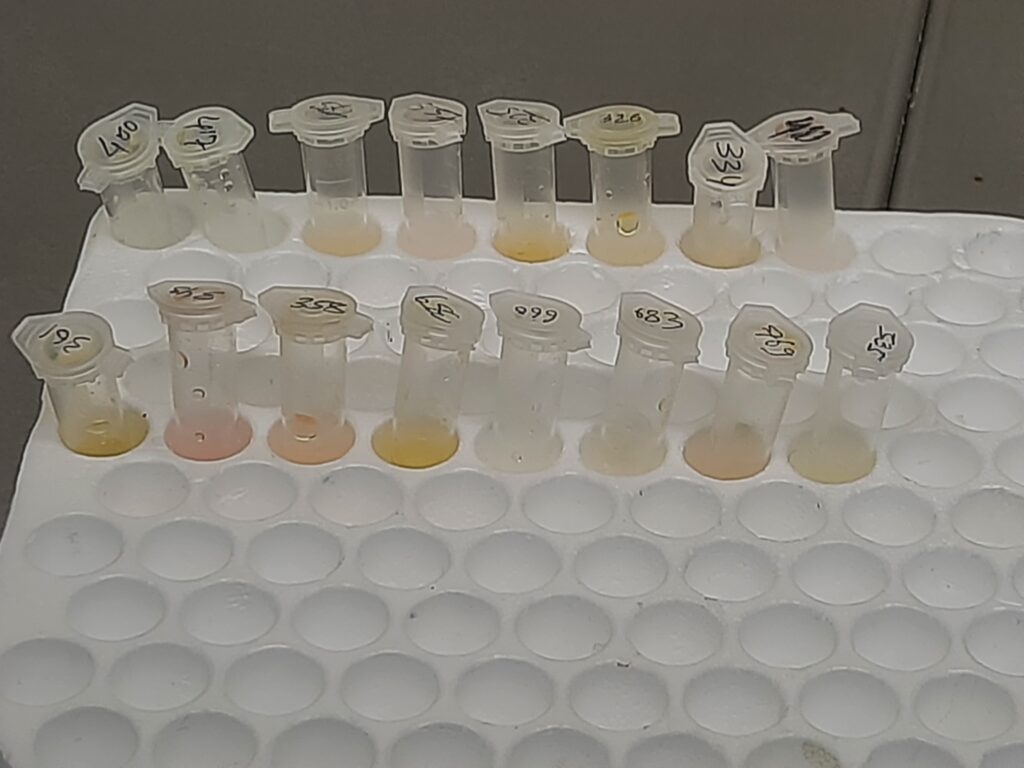
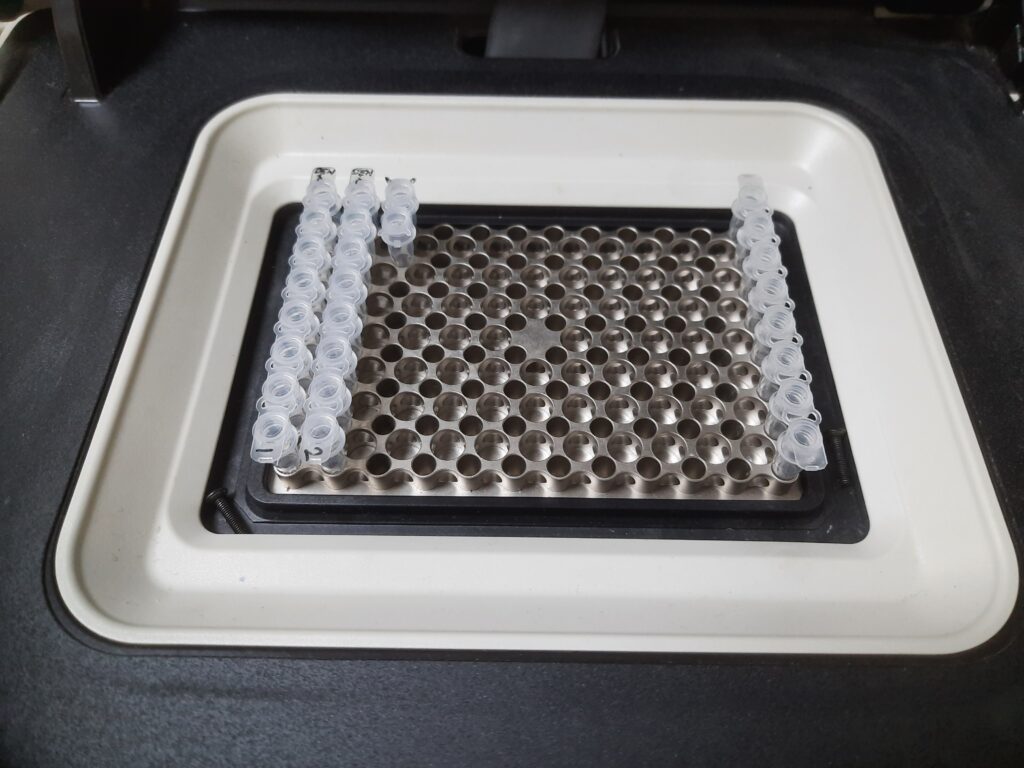





Result-Interpretation

First, observe the positive and negative control. If it is okay, go ahead with the tests. Interpret the result according given below table-
| S. No | Fluorochromes | Serotypes |
| 1. | FAM | Dengue serotype-1 (DENV 1) |
| 2. | Texas Red/Rox | Dengue serotype-2 (DENV 2) |
| 3. | HEX | Dengue serotype-3 (DENV 3) |
| 4. | Cy5 | Dengue serotype-4 (DENV 4) |
Keynotes on Dengue Serotyping Real-Time PCR Test
- The curve should be sigmoidal for result interpretation.
- More than one serotype may present in the same patient.
- Detection of Dengue virus serotyping infections is crucial for patient management decisions and epidemiological surveillance that shape public health policies.
Cut–off (CO ) Value or Reference Interval
Based on the clinical specimen test results the ROC (receiver operator characteristic) curve method is adopted to determine the CO (cut-off) value of these kits as follows.
- It is FAM positive if the FAM channel amplification plot shows an exponential increase with Ct <30.
- It is Texas Red positive if the Texas Red channel amplification plot an exponential increase with Ct<30.
- It is Hex positive if the Hex channel amplification plot an exponential increase with Ct<30.
- It is Cy5 positive if the Cy5 channel amplification plot an exponential increase with Ct<30.
RT-PCR Amplification
Put the complete PCR reaction tube into the fluorescent quantitative PCR analyzer, set the positive quality control detection hole, and set the sample name.
Total reaction system:30 µl
Fluorescence detection channel selection: FAM, Texas Red, HEX, and Cy5 channel are selected for detection: FAM is the serotype-1 indicator channel, Texas Red is the serotype-2 indicator channel, Hex is the serotype-3 indicator, and Cy5 is the serotype-4 indicator channel. The cycle parameter setting is shown in the table.
| Stage | Temperature | Duration | Cycle Number | Fluorescence Collection |
| 1. Reverse transcription | 52°C | 15 minutes | 1 | No |
| 2. Pre-degeneration | 95°C | 2.30 minutes | 1 | No |
| 3. Degeneration inside the cycle | 95°C | 15 seconds | No | |
| 4. Annealing and extension inside the cycle | 60°C | 30 seconds | 44 cycles | Yes |
Detection System Preparation
| Component Name | Dosage of a patient |
| Master Mix | 20μL |
| RNA of the specimen to be tested (template)/negative control/positive control | 10μL |
| Total volume | 30μL |

Thank you for sharing superb informations. Your site is so cool. I am impressed by the details that you have on this web site. It reveals how nicely you perceive this subject. Bookmarked this website page, will come back for extra articles. You, my pal, ROCK! I found simply the information I already searched everywhere and simply couldn’t come across. What an ideal web site.